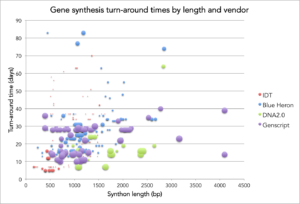
It’s been a while so I thought it was about time to do an updated post on observed turn times for commercial gene synthesis. After all, Rob Carlson posted updated cost and productivity curves for DNA sequencing and synthesis. As in my original post on this topic, I plot turn time vs length for orders placed at Ginkgo. The y-axis is turn time in days from when an order is initiated by Ginkgo to the day the company ships DNA back to us. The x-axis is the length of the synthetic DNA (synthon) in base pairs. Based on a request in the comments, I’ve also scaled the size of the data marker by how long ago the synthon order was placed/delivered to see if there are any observable trends in turn time over time (I don’t see any but it does show how we’ve tried out different providers over time 😉 ). The data has all the same caveats as last time – so for convenience, I’ve re-listed them at the bottom of this post.
I suspect that one of the most valuable aspects of this data, together with that from Rob, is that is shows how imperfect our benchmarks for the gene synthesis industry really are. For example, we don’t have a good metric for assessing companies on both cost and turn time. DNA2.0 has better turn times than Genscript but that comes at the expense of a 2X price premium – which company’s service is “better”? Selecting and then celebrating the right benchmarks is important because that’s where the industry will place its resources to improve the underlying technology. To date, the industry has largely been driven to reduce the per base pair cost of gene synthesis because cost per bp is the de factor comparable. This metric has pushed synthesis companies away from standardizing and competing on “library” offerings for sets of rationally designed synthons (many companies offer this but you have to get a custom quote which has a high transaction cost associated with it). Given where I suspect the engineering of organisms is going, making it even modestly more difficult to order libraries is probably detrimental to the field.
To help to combat this problem, I’d love to see the development of a true benchmark test for the gene synthesis industry – i.e. a set of synthons that spans different length, GC content, sequence complexity etc. gets designed and the orders placed simultaneously at all vendors so that there can be a true side by side comparison of the performance of all providers.
Probably the biggest update in the world of commercial gene synthesis over the last year and half is that multiple companies now have linear gene synthesis offerings. In no particular order, IDT offer gBlocks up to 750 bp for $139, Gen9 offers GeneBytes up to 3000 bb in length (they charge per bp but their rates aren’t posted online), and Life Technologies offer Strings up to 1000 bp in length for $149. With these offerings, gene synthesis companies have been finally able to break through the ~$0.30-$0.35/bp floor that they’ve been stagnating at from 2008-2012 by skipping the cloning and sequencing steps and shifting that technical risk onto their customers. The hope would be that these offerings could also result in lower turn times but jury’s still out.
Caveats to the data:
- Orders at the providers weren’t placed contemporaneously and we didn’t place orders for the same synthon at multiple providers.
- We design all our synthons using in house software – so the measured turn times reflect only ability of providers to build our synthons and not the quality of their gene design tools.
- By design, nearly all of our synthons pass all provider sequence checks and qualify as “low complexity” sequences from a synthesis standpoint.
- The synthons generally don’t result in protein expression which might adversely impact clonability by providers.
- To calculate turn time, I “start the clock” when we input the order to the provider website or send it to their sales rep and I “stop the clock” on the day the provider ships the gene back to me. So if there is a delay in order processing by the provider or rep, that counts against their turn time.
- There are a handful of synthons that providers couldn’t synthesize and/or clone. The 500 bp/80 day outlier for Blue Heron is one but there are also three others from IDT (unmarked). For those failures, I “stopped the clock” when the provider emailed me to say that they couldn’t make the synthon and weren’t going to try any longer.