From synthetic biology’s earliest days, DNA synthesis, and more specifically gene synthesis, has been touted as the central, enabling technology of the field. Gene synthesis is part of what lets us make the leap from the ad hoc, cut and paste of genetic engineering to the systematic design that is [or will be] the hallmark of synthetic biology. Given its central importance, it’s not surprising that many of us in the field keep a close eye on both gene synthesis technology and the gene synthesis industry as a whole. Yet most of the discussion focuses just on the cost of gene synthesis. Cost is important. But I’d argue that turn times are equally important in terms of how gene synthesis is used in the field – more on this below.
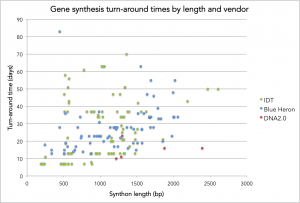
Below is a chart summarizing turn times versus length for DNA orders at Ginkgo. On the y-axis is the turn around time (in days) and on the x-axis is the length of the synthetic DNA (or synthon) in base pairs. [I refer to all of synthesized DNA fragments as synthons rather than genes since we don’t only synthesize genes.] Data points are colored by gene synthesis provider.
First, a few caveats:
- Orders at the three providers weren’t placed contemporaneously and we didn’t place orders for the same synthon at multiple providers. A more controlled experiment would of course be to place orders for a large set of identical synthons at multiple providers simultaneously and compare turn times. We didn’t do that.
- We design all our synthons using in house software – so the measured turn times reflect only ability of providers to build our synthons and not the quality of their gene design tools.
- By design, nearly all of our synthons pass all provider sequence checks and qualify as “low complexity” sequences from a synthesis standpoint.
- Almost none of the cloned synthons result in protein expression which might adversely impact clonability by providers.
- To calculate turn time, I “start the clock” when we input the order to the provider website or send it to their sales rep and I “stop the clock” on the day the provider ships the gene back to me. So if there is a delay in order processing by the provider or rep, that counts against their turn time.
- We rarely pay the 2X+ price premium to synthesize genes at DNA2.0 – hence the limited number of datapoints for them.
- There are a handful of synthons that providers couldn’t synthesize and/or clone. The 500 bp/80 day outlier for Blue Heron is one but there are also three others from IDT (unmarked). For those failures, I “stopped the clock” when the provider emailed me to say that they couldn’t make the synthon and weren’t going to try any longer.
OK caveats finished. What can we infer from this chart? Here are two takeaways.
Lesson 1: Gene synthesis can’t be a part of the design-build-test loop until turn times improve dramatically
Based on our data, turn times are highly variable and show little to no correlation with length overall. This means that when you place an order, you have no idea if you’ll get it back in 2 weeks or 5 weeks. From an engineering process standpoint, I’d argue that the unpredictably long turn times mean that it is crazy to include outsourced commercial gene synthesis in the design-build-test loop as you try to engineer an organism. Instead, do gene synthesis orders up front as a batch (thereby hopefully eliminating gene synthesis from your cycle time) and then mix and match the synthesized parts via a DNA assembly technology with a faster turn time. Or alternatively, try to achieve faster turn times by doing gene synthesis in house from oligos.
Lesson #2: Different providers specialize at different orders
IDT appears to be quite fast at making sub-500bp synthons. This is not too surprising given that IDT leverages their ultramer oligo synthesis tech to offer flat rate pricing on so-called minigenes (< 400bp synthons). At that length scale, you can also opt to stitch together oligos to make the part yourself. But between the costs of oligos, cloning reagents, sequencing and your own time, you might not do much better than the cost of a minigene (even factoring in cheap grad student/postdoc labor!). For synthons in the 500-1500 bp range, Blue Heron seems to be a reasonable compromise choice in terms of turn times versus costs. You get industry competitive pricing with decent turn times. Overall, DNA2.0 appears to have the best turn times for >1 kb synthons. Admittedly this is based on a very limited sample size but anecdotal rumors from folks in the field back it up. So if you’re in a rush and can tolerate the 2X price difference, DNA2.0 could be the way to go.
I’ll close by saying that this post is in no way an attempt to rag on gene synthesis providers. Building DNA is tough. And building DNA for customers is even tougher. But it’s important to think hard about what the realities of costs and turn times of commercial gene synthesis mean for developing best practices for engineering organisms going forward.