In our last blog post, Vichka and Etowah told us about the cool things that they did this summer. This week, Liam and Aileen tell us about their internships on the Software Team!
Liam Bai, Software Engineering
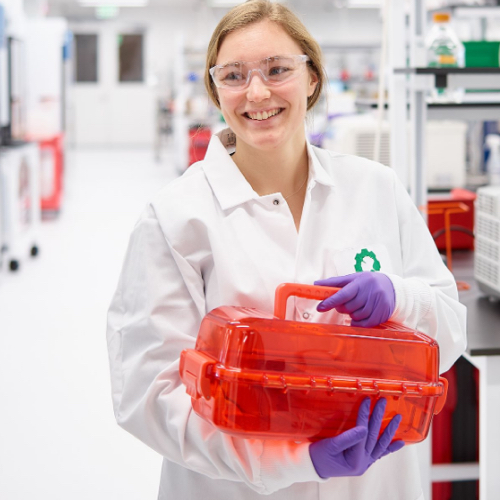
Hi! My name is Liam, and I study Applied Math and Computer Science at Brown University. This summer, I interned with the Base Chasers team (each of the scrum teams at Ginkgo has given itself a fun name), and worked on expanding Ginkgo’s next generation sequencing (NGS) data pipelines. Specifically, I built a pipeline orchestrating sequence data from Oxford Nanopore long-read sequencers.
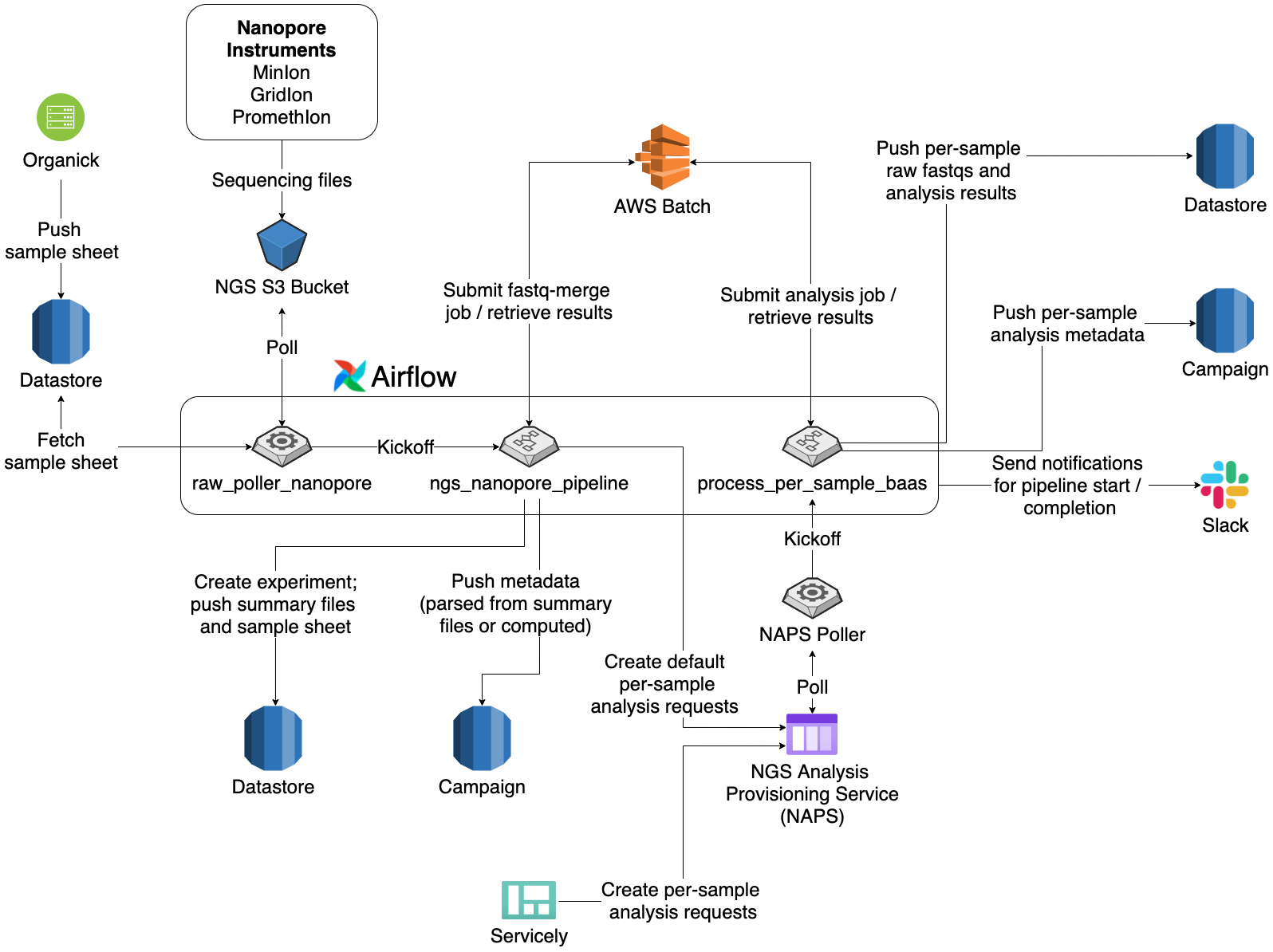
Although Nanopore sequencers are not new at Ginkgo, software support for the instruments was minimal. An internal command-line tool handled most of the data processing, and any downstream quality control and analyses were done manually by bioinformaticians in Jupyter notebooks. The goal of my project is to replace the current processes with a pipeline that is more automated, scalable, observable, and robust, with additional features including metadata capture, notifications, and support of custom analyses. In short, after starting a sequencing run, a sequencer operator can sit back and relax, knowing that data––along with metadata such as QC statistics––will show up in the right places in the right format.
The Nanopore pipeline runs on Airflow, an open-source workflow orchestration system. The pipeline integrates with Datastore and Campaign––internal data/metadata storage services at Ginkgo––along with the NGS Analysis Provisioning Service (NAPS), an internal queuing service for analyses. To improve efficiency and scalability, I used AWS Batch to process large, raw files, compute metadata, and run analyses.
Zooming out, as part of the testing pipeline that allows scientists to gain detailed insight into the strains they work with, NGS plays a critical role in Ginkgo’s mission to make biology easier to engineer. Newer long-read (Nanopore) sequencers complement short-read (Illumina) workflows and enhance our confidence in the sequence data. It was immensely satisfying to see my project contribute to Ginkgo’s efforts in evangelizing standardization and building out infrastructure that can support engineering biology at this unprecedented scale.
I loved being a part of the Base Chasers this summer, and learned a ton from their mentorship. Perhaps more importantly than learning the ins and outs of Airflow, I picked up on many design patterns that make a system robust and scalable, and learned the importance of communication in building software.
I am extremely lucky in being able to come into the office for the latter half of my internship. I bonded with my teammates and fellow interns, and loved the culture of whimsy at Ginkgo. Every day, I am inspired by the Bilobans’ passion for making biology easier to engineer, while constantly reminded that there is so much fun to be had along the way.
Aileen Ma, Software Engineering
Hi! My name is Aileen. I’m a rising senior at MIT majoring in 6-3 (computer science) and a full stack software intern on the Terminators team for the summer of 2021. The Terminators team primarily works on Loom, an internal platform for designing and ordering DNA constructs. I was remote for the first half of my 11-week internship, but returned to Boston for the remainder of the time.
While working at a synthetic biology company in the midst of a global pandemic is already a one-in-a-million experience, going public also adds a unique dimension to my time here. I have really enjoyed not only learning about producing scalable technology, but also in watching the steps a startup takes as it rockets into the public eye.
My Projects
My two projects focused on improving the user experience with Loom. There were two main aspects of Loom that I worked on: the search feature and the bulk editing feature. As a full stack intern, my two projects were chosen to give me experience with both the frontend and backend. Improving search was a project largely concentrated on the backend, while bulk editing was a combination of both, although a little heavier on the frontend.
Making Search in Loom Delightful
The Loom search feature is used to look through design units and taxa. Design units are the building blocks used to create DNA designs. Taxa refers to the organism species that produces a sequence. The previous implementation used matching by trigram similarity, a text searching algorithm supported by Django which ranks results based on matches with the search query by checking every three characters (a trigram) and filtering only those that meet a certain threshold of similarity. In practice, there were many common search queries that would not meet the similarity threshold even though the user intended them to. For example, if the user searched “large table”, the search result “large brown table” may not meet the similarity threshold. Additionally, this method is very slow because it would be necessary to loop through every trigram and compare with the existing entries, and users would often rather select a pre-loaded default than wait 20 seconds or so to search for the proper taxon (if it would even appear at all).
Because the taxon database is fairly constant while the design units and design databases are frequently being updated, it is best to use two separate methods of searching through them. For the design/design units databases, it turns out that a more accurate method involves regex matching for every word in the search query (a Postgres LIKE query) against each entry within the desired fields. This seems a little brute force, but proved to be twice as fast as trigram search and better supported user intentions.
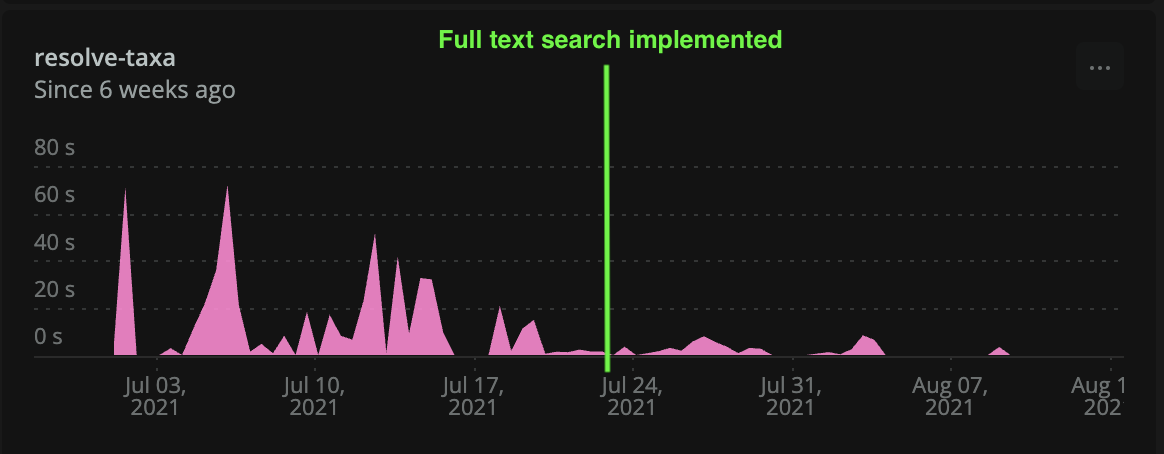
The taxon database is much larger than the design unit database, but it updates less frequently. Instead of using a brute force method similar to the one used for designs/design units, it was much more efficient to implement a search vector with GIN (generalized inverted index) for speedy lookups. GIN has a higher build cost than the previous method (GIST), but faster lookup times. For a database that doesn’t change very much (and doesn’t need to be built frequently), GIN is the way to go. Results were between 2x to 6x faster than before, along with better accuracy. Results were also limited to the top 100 matches, which helped speed up the display dramatically.
Editing Feature
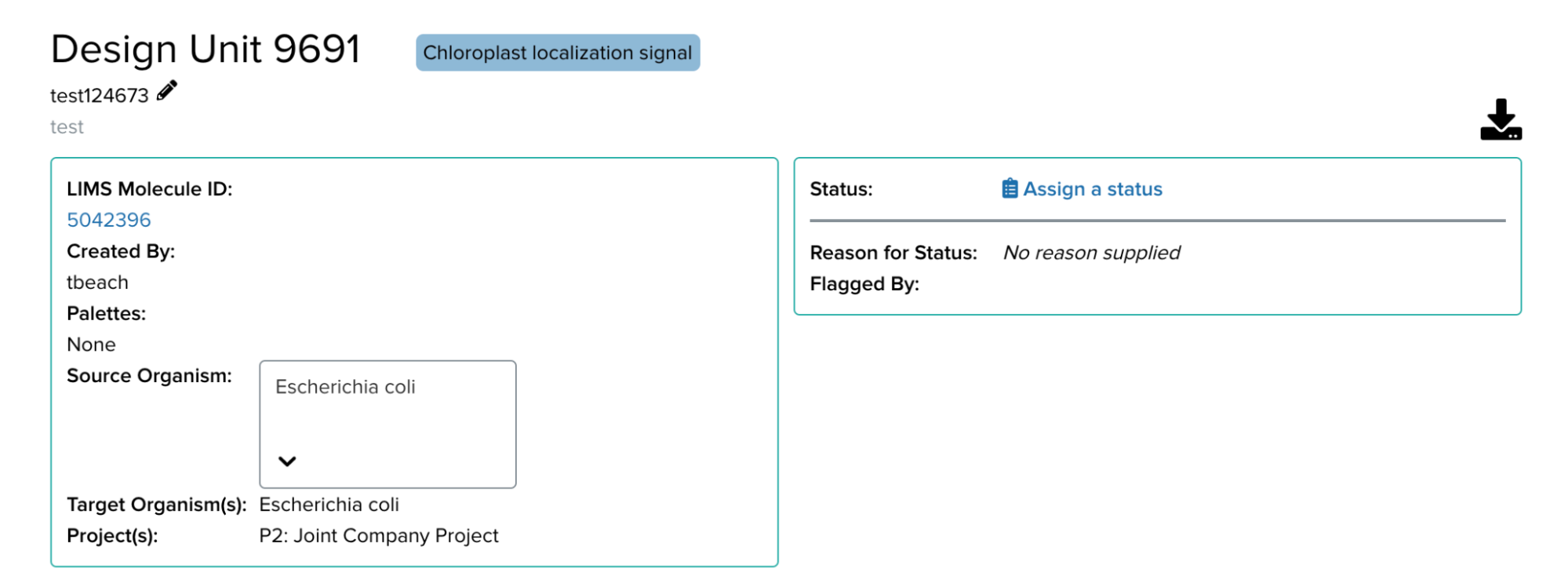
Once a design unit is created, it can be difficult to change the metadata associated with it. A user who may have hastily selected the default E. Coli as the source taxon may realize they want to change it later on (especially now that they can quickly find the appropriate taxon!). Providing an editing feature is therefore very desirable to enhance the user experience. The editing feature came in two parts: adding more features to single unit editing, and adding a bulk editing feature.
Previously, design units could be modified by name, description, or status. I worked on expanding upon these features so that design units could also be modified by source taxon, target taxon, project, or part types (which are used for characterizing and grouping design units) for a single design unit. This involved adding an appropriate addition to the back end that would allow for these new mutations and writing unit test cases. Throughout the process, I became familiar with GraphiQL for making queries – this allowed me to figure out if mistakes were happening on the back end or front end. On the front end, I worked on integrating editable features with existing components such as dropdown menus. React is a great framework that allows for components to be reused from various parts of the platform, allowing for very scalable software.
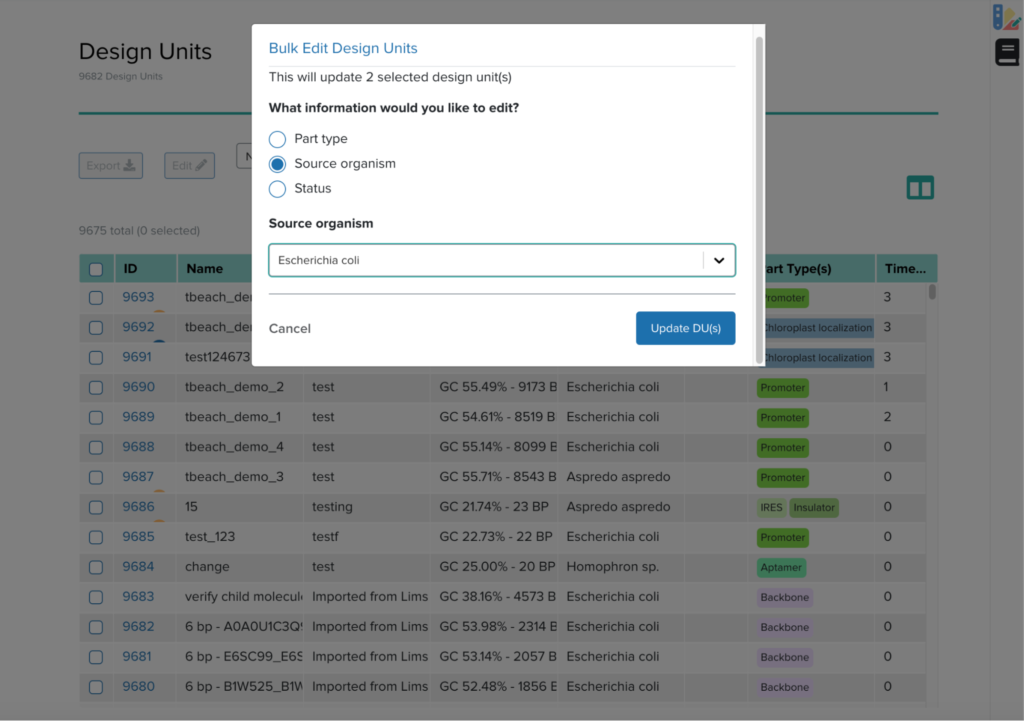
Finally, I also worked on the bulk editing feature, which will override a single field with a user input. As Ginkgo grows larger, the internal database of design units grows increasingly large. With a bulk upload feature integrated with the existing software, it becomes important to easily fix small errors in many different entries. Bulk editing seeks to implement this feature. I worked with my mentor and the Lead UX Designer to figure out how the user should interact with bulk editing. Similar to my experience with implementing a single edit feature, I started with adding relevant features in the back end and moving towards the front end. The end result was a lovely modal as shown.
Social Experience
Ginkgo hosted 11 interns this summer: 6 software interns, 1 product manager intern, and 4 business and development interns. One of the benefits of having a smaller intern class is forming a very tight-knit community. The onsite software interns were particularly close, given that we were all around the same year (rising college seniors/freshly graduated). We spent a lot of time together, both in and out of the office. One time, four of us even stayed in the office until midnight discussing a combinatorics question proposed in the #help-science slack channel. Sometimes when we leave early, we’ve eaten out at various restaurants in the area, hopping from taiyaki ice cream to pizza to fried chicken.

Ginkgo has also hosted a few intern events, the most prominent of which was the catered lunch with our founders. They answered every question we threw at them with complete transparency. In fact, the whole company is pretty rooted in transparency – documents are easily accessible to employees, including meeting notes, project documentation, and OKRs. The company also hosted an intern/mentor dinner at Committee, where we completely stuffed ourselves with Mediterranean food and got the chance to speak with other interns/full time employees we didn’t typically interact with.

After the intern/mentor dinner, I also got acquainted with some of the business interns and learned more about their projects. They are all in the midst of pursuing their MBA degrees and have been great about reaching out to the software interns. They have such vastly different experiences from us, having already accumulated some experience in the workforce, and it’s fascinating to hear about the path that brought them to Ginkgo.
On Friday nights, there would often be happy hour or other social events happening in the kitchen after a long week of work. Chess and other board games are popular pastimes, and I met many other people at the company through these. Bughouse (2v2 chess) is a popular variation here and draws a bit of a crowd.
Interacting with the Company
I met daily with my mentor, usually on-site, since I tend to ask questions that are more easily answered in person. Most of my team was remote, although some of them worked in the area and would come into the office occasionally. Every two weeks, we would have sprint planning sessions that allowed me to interact more with the team; otherwise, I usually spent most of my time with my mentor. I worked very closely with my mentor, and as someone who was much more familiar with the codebase and tools for working with software, he guided me through many patches of my internship where I needed help. I learned so much from seeing the way he thought about problems and digging to the root of the issue that by the end of my internship I could solve problems about 5x faster than I was at the beginning of my internship.
One of the best parts of being an intern is being able to reach out and ask questions without feeling awkward about it. We had weekly AMAs with a different Digital Tech Team member every week, and it was extremely insightful to chat with them about their experiences and backgrounds. We heard from solution engineers, software architects, and software engineers on different teams. Many of these people previously came from other companies working in healthcare or biotech, and knew each other prior to joining Ginkgo. One of the questions that seemed to garner a mixed variety of answers from people was the path of either developing a broad set of skills or a very deep understanding of one particular field. As a biology company that seeks to sell a service, I originally imagined that having a solid foundation in both biology and computer science would be helpful. While the software engineers all have an interest in biology, biology background is not critical. This question of pursuing a broad versus deep set of skills is answered individually andI look forward to exploring further myself.
Aside from AMAs for the interns, many of the groups here hold office hours to explain the projects they’re working on. For software engineers, office hours are a fantastic way to learn more about the biology side (and vice versa!)
While we technically have a hierarchy, the organization feels very flat. As an intern group, we’ve spoken with every level of the organization up to the founders, and even cornered Tom Knight himself to ask questions about the founding days. Our head of software is very involved with the internship program, and sometimes joins us for lunch or happy hour chats. The happy hour events have also been very fun and attended by a variety of people across the company, and provides another avenue to learn more from others.
Concluding Thoughts
These past 11 weeks have flown by, especially being in person. I had a wonderful time learning new technology, speaking with fascinating people, and working on a product that will make a difference for others. As Ginkgo scales up, there will be more and more people relying on our internal software and it’s been important for me to remember that good software practices now will make life exponentially easier for future maintainers. There is a strong company culture here that incentivizes growth, with the atmosphere of a startup but enough resources to make the Batcave seem paltry in comparison. People here are truly passionate about shaping a future with synthetic biology that feels like it’s just around the corner, and working here has been an optimistic reminder of the achievements we might have in the years ahead as we continue down this path.
We’ll hear from Kevin and Vidya next, so check back soon for that!
(Feature photo by frank mckenna on Unsplash)
