In our last blog post, Ariel, Yasmine and Evan told us about their summer. Lets hear what Delight and Judah have been up to!
Delight N., Product Management
Hi! My name is Delight and I’m currently studying bioengineering at MIT. This summer, I interned on the Product Management team at Ginkgo, working on the development of a catalog for Ginkgo’s Foundry offerings. The Foundry refers to Ginkgo’s lab space and teams that each specialize in a different area, such as DNA design or high throughput screening. “Offering” is a term used to describe the protocols or work that Foundry teams do for other teams. Ginkgo’s Foundry currently supports over 100 offerings with a variety of uses and complexities. As Ginkgo continues to grow rapidly, understanding what the Foundry can offer is key to leveraging Ginkgo’s resources most effectively. This is especially important as the company continues to scale because if people are not aware of what offerings exist, they cannot take advantage of them.
To achieve this end, we proposed an offering catalog that organizes key service information. The guiding belief was that by improving presentation, accessibility, and information about Foundry offerings, users could more easily identify and request Foundry offerings. The goal was to provide a single source of truth for Foundry offerings that allows for browsability by scientific application and discovery of new offerings.
Understanding the needs of the core users was an important first step to working towards a solution. This included understanding information they needed access to, what functionality helped them navigate with ease, and what styles of presentation were intuitive and easily comprehensible to them. The target users of this catalog are Ginkgo’s organism engineers and project leads who can leverage Foundry’s tools and processes to fulfill the needs of Ginkgo’s customers.
After better understanding the value users needed to extract from the catalog, I worked with Darek Bittner, a seasoned product designer to design the general layout of a catalog. To inform our brainstorming session, I researched presentation styles of product libraries from a variety of industries, paying special attention to elements that helped these libraries convey key pieces of information to readers. Based on the information gathered I produced thumbnail sketches, some of which are shown below.


After aligning on key elements for the catalog, the next step was to develop a prototype. We decided to use Tableau, which is an interactive data visualization software, because it would allow us to quickly iterate on the prototype without software engineering support. Darek and I created three different versions prior to the first round of user testing. We then conducted user research using the prototype where we assessed how intuitive, easy to use and flexible our design was. Three of our most important findings were:
- Users were able to find offerings based on their name, description and intended purpose
- Descriptive names helped users navigate through the catalog and understand the nature of Foundry offerings
- Using tableau as a platform resulted in usability issues that could pose significant challenges to implementing it as a long term solution. We also found that users chose to find offerings by searching through Foundry teams, rather than via scientific categorization, which was unexpected.
We used these findings to inform the updated design of our prototype in Tableau, thus resulting in the current version of the platform, a screenshot of which is shown below.
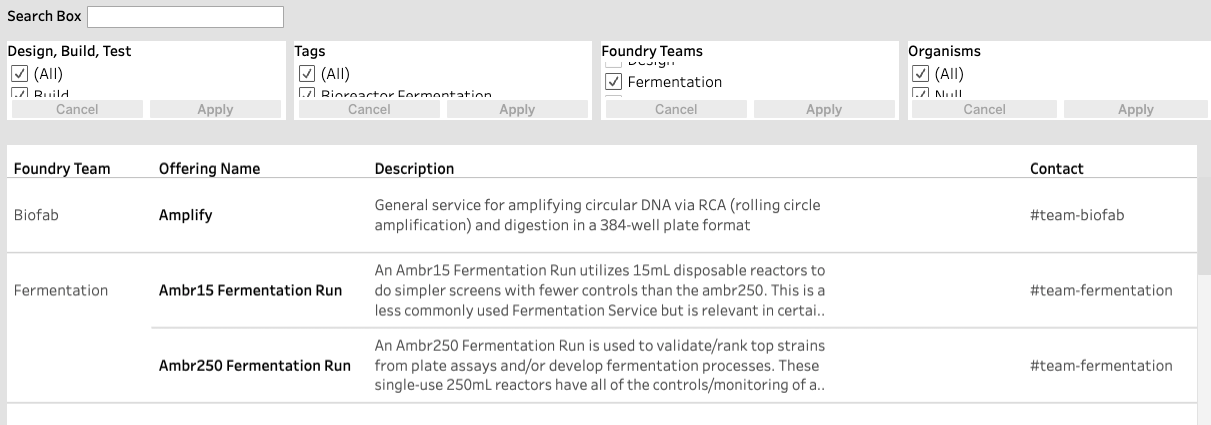
In order to improve search and filtering for our users, we decided that we needed to provide additional guidance to the Foundry representatives who directly contribute to the catalog on how information should be presented. Based on this, a guideline was created requesting that representatives select all valid organisms for each offering.
To address the usability issues caused by Tableau, I looked into alternative platforms that could be used for prototyping and possibly support the minimal viable product for the system. In the interim, we will release the prototype we created because, with its basic functionality, it will provide tremendous value to users. In the future, we would like a more stable version of the catalog to contain information that is easy to understand and navigate by, be updated automatically based on the information entered into Smartsheets, and work in sync with the offering creation tool in Servicely, which is another Ginkgo software product.
Working on this project this summer has been an interesting introduction to software-oriented design and has given me a great opportunity to practice design thinking. I learned the importance of paying attention to details and evaluating how they affect the outcome of different solutions, while keeping in mind the big picture. Keeping an open mind was central to moving forward after discovering ways that the design was interfering with our users’ ability to use our prototype. Our solution went through several versions, most quite different from the last, but by combining and altering the presentation and function of different elements, we discovered new ways to design our solution and gained valuable information on how best to support our users.
Judah Wyllie, Software Engineering
Introduction
Hi! My name is Judah Wyllie, and I’m a recent graduate from the University of Washington with a Bachelor’s degree in Computer Science. I had the amazing opportunity to be an intern on the Infragon team at Ginkgo this summer. The Infragon team is a hybrid software/automation team that works at the intersection of biology, software, and automation.
Main Project: SCLE Schedule Cache
Background
At Ginkgo, scientists use Autoprotocol to define biological protocols that are executed on integrated sets of lab automation devices called workcells. At a high level, a protocol consists of a set of containers and a sequence of instructions to perform on them (e.g. spin, seal, incubate). Before a protocol can be executed, it must be “scheduled” using a tool called SCLE, which outputs low-level instructions for the workcells to execute.
Problem Statement
Once a protocol is validated (i.e. it performs the scientific intent correctly/reliably when executed), it is often operationalized as a Foundry Service and is run repeatedly for weeks, months, or even years.
This can be challenging however due to a number of intrinsic limitations of the SCLE scheduler. First, scheduling large protocols can be extremely time consuming – several hours in the worst cases. Second, scheduling is non-deterministic, meaning that scheduling the same protocol multiple times can yield different results each time.
The goal of my project was to address these limitations by designing and implementing a caching service that allows for the reuse of previously computed schedules for a given protocol, thereby reducing scheduling times from hours to seconds, and increasing the reliability and reproducibility of the scheduler.
At a Glance
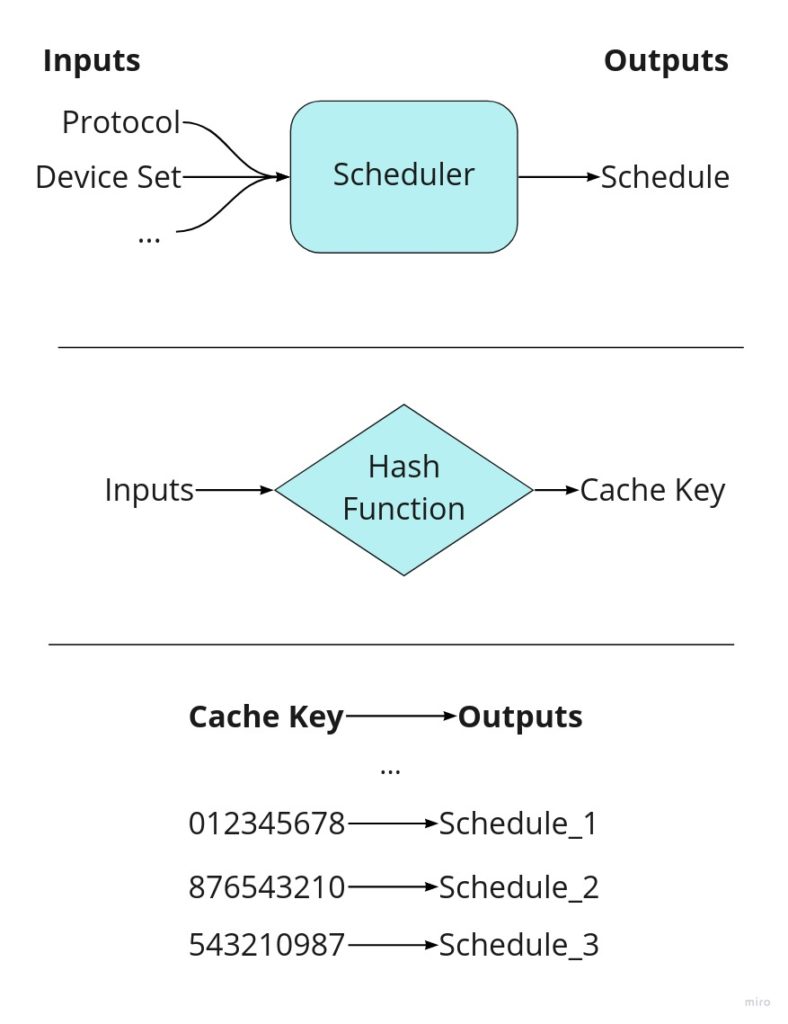
At a high level, the cache service works by storing scheduling outputs and reusing them when given the same inputs, thus bypassing scheduling altogether. We can tell if our schedule is stored in the cache by use of a cache key: a list of inputs that identifies unique results. The cache key is composed of a MD5 hash of the provided workflow as well as other unique identifiers. The logic behind the service’s decision making is simple and can be followed from the diagram below.
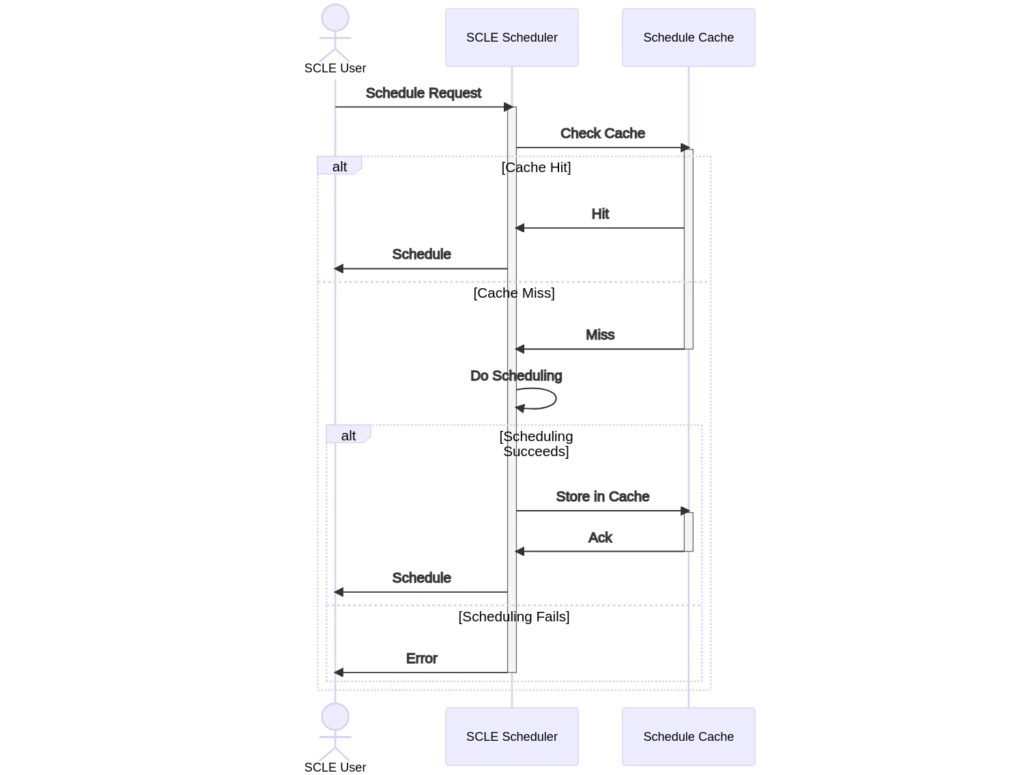
The SCLE User requests the scheduling service for a given protocol, plus other scheduling inputs. The cache service then checks the cache to see if there is a match. If there is a hit, it can simply return the cache object that stores a compatible schedule. In the event of a miss, the scheduling service continues as usual. If the scheduling job is successful, it will then store the result in the cache for future usage.
Solution Implementation
Now we have the core functionality of our service in place, but the bulk of my work during my internship actually revolved around building a robust and reliable service with all the features you’d expect from a modern API. Here are some of the main decisions we made:
- Our API is built on FastAPI, a Python framework with built-in type validation and Swagger documentation generation.
- Our service is containerized within Docker, which allows us to easily test it locally in a contained environment.
- An AWS Lambda hosts our service (through a Docker container). This “serverless” platform allows us to scale easily. Our service is easily deployed to our Lambda via a CI/CD pipeline.
- Cache objects are stored in a PostgreSQL database hosted by Amazon RDS.
- Unit tests and integration tests are integral to a service like this, where errors could have real-world consequences. Our test suite automatically builds a new database to run its tests on that matches the production environment. Real-world schedules and workflows are used in the test suite.
- Metrics and errors are reported to New Relic. A custom dashboard allows us to view metrics like Cache Hit Rate and Total Time Saved.
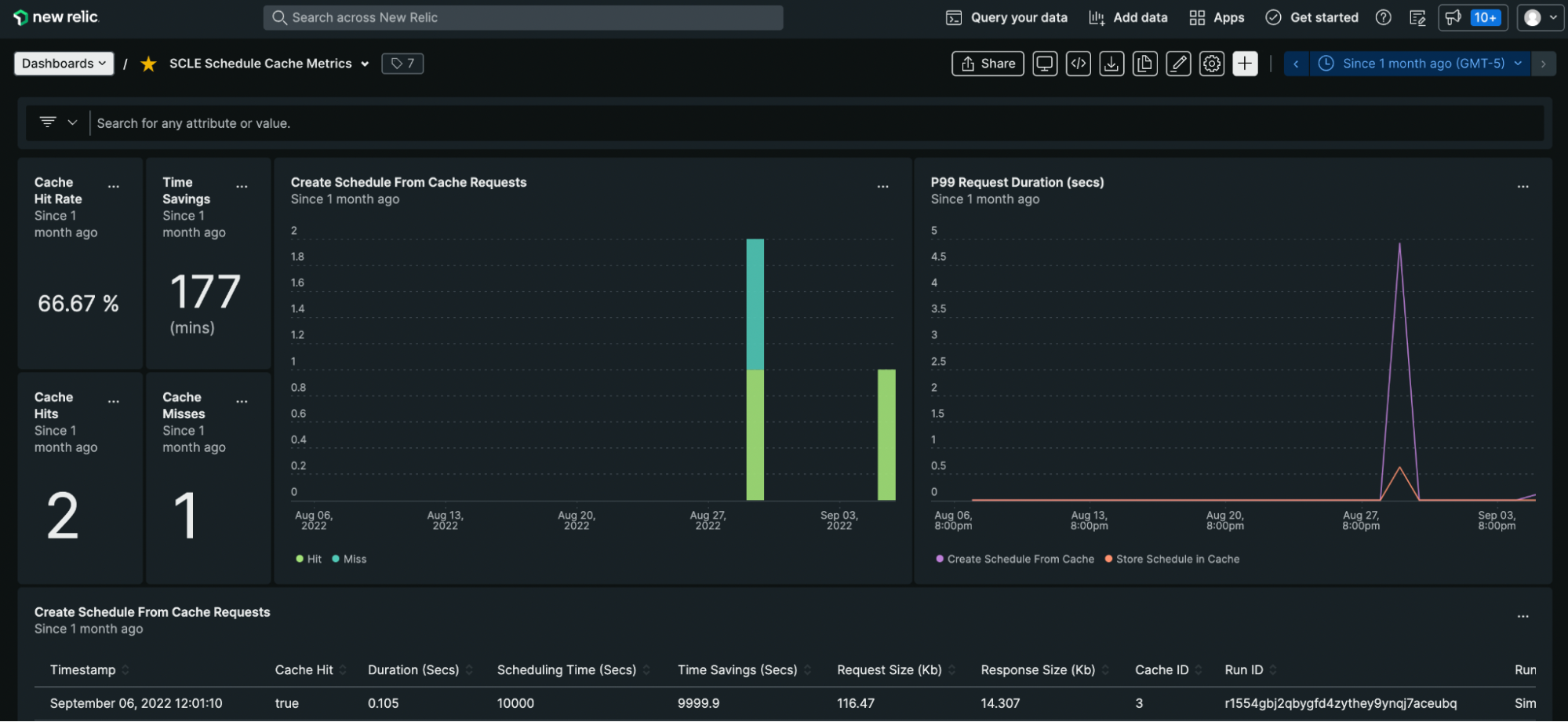
My Ginkgo Experience
This summer marked my first time on the East Coast, as well as my first introduction to biotech. Ginkgo was a great place to experience both of these things! I especially valued the experience of going into the office and being able to see firsthand the work being done in the labs. It was also a great opportunity to meet new people and talk to people who worked in all sorts of roles. Being able to actually talk face to face with the people you work with was such a stark contrast to the remote internships and remote learning from the past 2+ years. Ginkgo also allowed me the flexibility to work from home, which was a nice perk.
Some highlights from my summer include meeting the founders, weekly intern lunches, intern events like Canobie Lake Park and kayaking in the Charles River, and exploring the Boston area on my own.
Conclusion
As this summer wraps up, we want to thank our Digital Tech Team interns for their contributions. What they accomplished has been impressive. Thank you, and we wish you great success in what you all will be doing next!
(Feature photo by ian dooley on Unsplash)
