Biology by design.
Biology is the most advanced manufacturing technology on the planet. We program cells to make everything from food to materials to therapeutics.
What We Do
End-to-End Capabilities. Unparalleled Scale.
Explore Ginkgo’s capabilities for therapeutics and vaccines, agriculture, nutrition and wellness, and more.
Enzyme Services
Discover, engineer, optimize, and scale up enzymes for your application
Protein Services
Discover and optimize the production of novel proteins for a variety of applications
Metabolic Engineering Services
Engineer and optimize biosynthetic pathways for small molecules
Strain Optimization Services
Screen millions of strain variants to optimize for your application
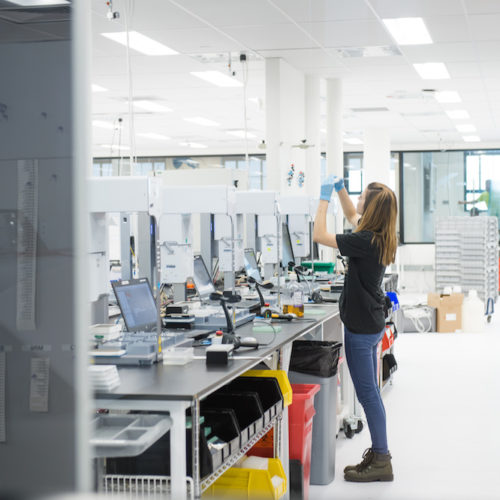
Process Design & Scale Up Services
Scale up your fermentation from 250 mL to >50,000 L on our platform
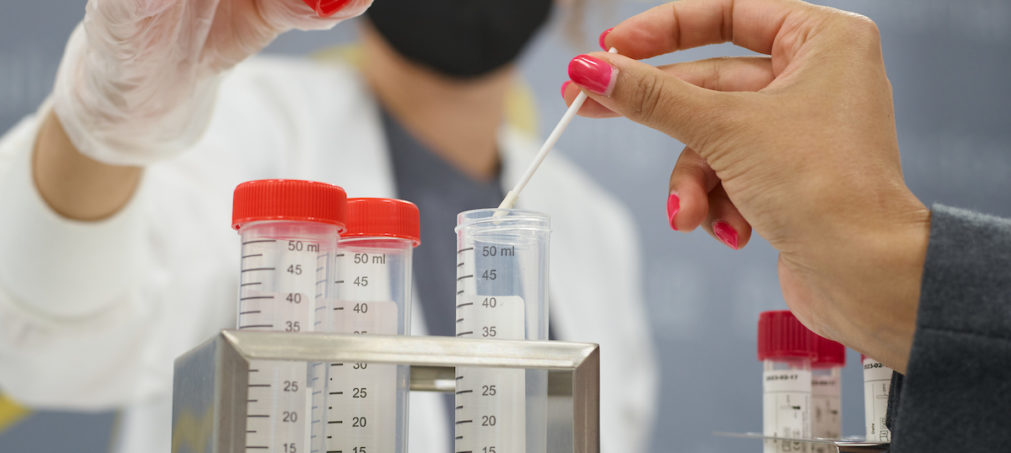
Biosecurity Services
Design a multi-layered biosecurity program to protect the things you care about the most
Our Projects
Markets We Serve
Explore how biology is being used across industries and how Ginkgo can help you.
Agriculture
Crop protection and nutrition, animal health and wellness, carbon solutions: we’re here to help
Biopharmaceutical
Use our synthetic biology platform to discover, optimize, and scale up a wide range of therapeutic modalities
Governments
Secure the things you care about the most — your people, your environment, your home
Industrials
Chemicals, materials, fuels, waste, mining, water treatment: biology is revolutionizing industrial processes
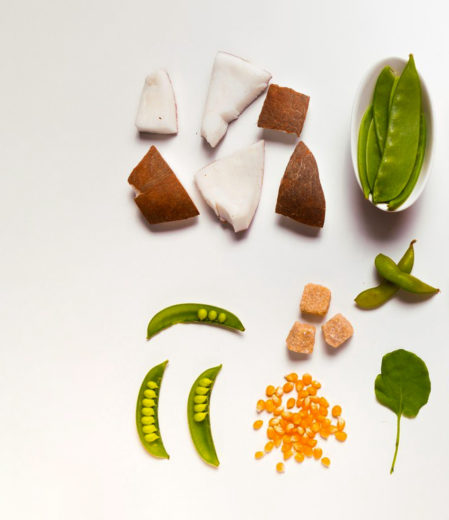
Nutrition & Wellness
Improve proteins and enzymes, fats and oils, flavors and fragrances, and so much more with synthetic biology
…and many more!
Speak with the Ginkgo team today about your industry’s needs and how biology can help solve them
Making biology
easier to engineer.
Technology