In late March, a group of researchers at the University of California, San Francisco published a preprint describing a protein-protein interaction map that could find potential drug targets against SARS-CoV-2. Their study looked for interactions between every viral protein and every human protein, with the idea that molecules known to bind those human proteins might disrupt the interaction with the viral protein, therefore interfering with the viral life cycle. Several hundred hits were identified, and of those hits, over 50 protein targets had known small molecule binding partners, so-called “druggable” targets. These small molecules could potentially interfere with viral protein binding and therefore be repurposed against COVID-19. One interaction was observed between the viral protein Nsp13 and human centrosomal protein 250 (CEP250), a protein target previously predicted to be too flat for small molecule drugs to bind to it and therefore “undruggable”.
Several years ago, our team at Warp Drive Bio performed a very different high throughput screen, searching for human proteins that could bind to a novel natural product that the team had discovered. The molecule, called WDB002, is made by a species of soil-dwelling Streptomyces bacteria that was identified using Warp Drive Bio’s genome mining technology. WDB002 was found to be a strong binder to a little appreciated human protein called CEP250 — the same protein identified in the recent SARS-CoV-2 study! WDB002 first binds to another human protein, FKBP12, which then forms a tight three-part complex with a portion of CEP250.
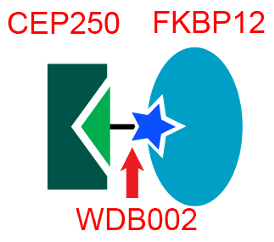
In October 2018, Revolution Medicines bought Warp Drive Bio, and Ginkgo then acquired the Warp Drive genome mining platform from Revolution Medicines in January 2019. Today PNAS published an article authored by members of the former Warp Drive Bio team, showing how WBD002 and its related molecules can bind to the “coiled-coil” domain in CEP250, in complex with FKBP12 via binding studies and crystal structures. These coiled-coil domains are very “flat” and therefore notoriously difficult targets for drug discovery, which makes the tri-complex modality potentially useful for finding leads against other so-called “undruggable” disease targets.
As for CEP250’s role in COVID-19, we don’t know yet whether WDB002 will be useful for treating the virus, but Ginkgo is committed to finding out. Revolution Medicines has licensed intellectual property to Ginkgo to answer this question. Our team has already produced a batch of pure WDB002 from Streptomyces. Now, we are testing how it behaves in an in vitro live virus assay in collaboration with research groups that can perform these studies at biosafety level 3. It’s been exciting to be part of the story of WDB002, from when it was first discovered to exploring its potential as an antiviral. Despite the unknowns inherent in drug discovery and development, it is inspiring to witness Ginkgo’s ability to apply its platform towards vaccines, therapeutics, and diagnostics for COVID-19. We hope to share news about the WDB002 project in the coming months.
