Ginkgo’s biological foundries are automating and scaling the process of organism engineering, allowing biological engineers to prototype thousands of biological designs. Our Foundry is organized as a number of services that receive and operate on requests from our organism engineers for designing, building, and testing organisms at a massive scale. These service requests are submitted via Ginkgo’s Software Portal to a foundry service software platform that processes those requests to perform biological computations, generate and run robotic workcells, and return the data generated back to the user.
Current Architecture
When Foundry services were introduced into the new Ginkgo Portal software in 2019 we decided to start with the NGS (Next Generation Sequencing) and HTS (High Throughput Screening) services. The goal was to build a software architecture to successfully handle service request submission, asynchronous processing of the request, and data return. Like we did with many other Ginkgo software applications at the time, we decided to go with Django as a server, and Celery for asynchronous tasks. The rollout of NGS and HTS services was a huge success and the architecture met our needs at the time.
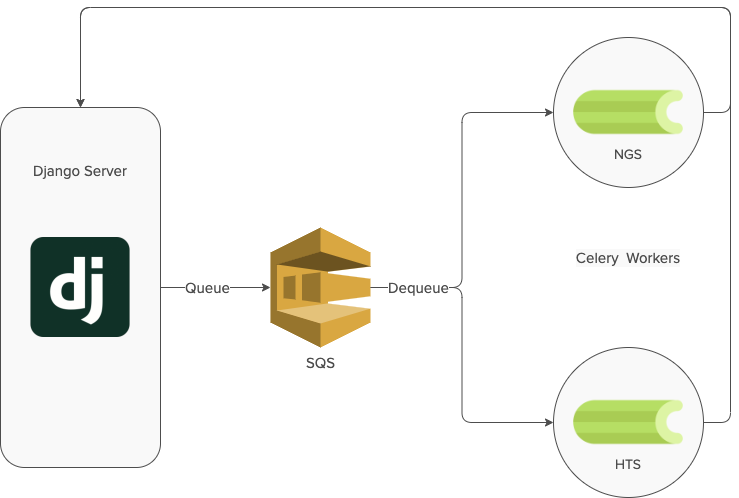
As we onboarded tons of new foundry services in 2020, we realized two main problems with the Celery architecture.
Scalability
One of the key design decisions we made was that every foundry service was run in an independent Celery worker container. This allowed achieving upkeep of one service without affecting the others, and also prevented bugs introduced from one service into all the other services that were running. However this meant the number of containers running Celery tasks increased every time a new foundry service was added, making it more difficult to manage the configuration and maintenance of the new containers. There was also a high likelihood of Python package incompatibility between services.
Availability
Many of the foundry services run asynchronous tasks that can run for several minutes. This makes it difficult to bring the service down during deployment of the new code, and drain time isn’t always reliable.
New Architecture
To solve the scalability and availability problems we decided to explore serverless architecture in place of Celery. In an ideal serverless world we would:
- Easily scale foundry services without bringing up new servers and containers.
- Deploy new code without terminating anything running on the old code.
- Run each foundry services independently without affecting each other.
- Decrease cost by having foundry services up only during the course of processing requests rather than 24/7.
Most of the code we had was already running on AWS, making Lambda a great serverless option. But because of in-house expertise we also wanted to keep using Django in the new architecture.
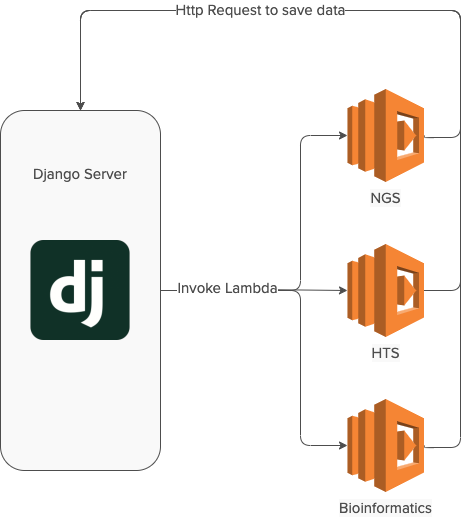
We wanted to explore serverless architecture as shown in the diagram above to scale foundry services.
Why Zappa?
After doing some research we found Zappa, a Python library that makes it super easy to build and deploy event-driven and server-less Python WSGI web apps. It has an awesome feature in that it wraps a Django application as a Lambda which is deployed to and hosted on AWS. Upon further examination of Zappa documentation it was impressive how little setup and few commands were needed to have an application fully up and running. Furthermore it came with many features such as free SSL certificates, global app deployment, API access management, automatic security policy generation, precompiled C-extensions, auto keep-warms, oversized Lambda packages etc.
Our familiarity with Django apps and the many cool features it provided made Zappa a very compelling option. Hence we decided to use Zappa to build out and evaluate a prototype of the proposed serverless architecture.
The Zappa Prototype
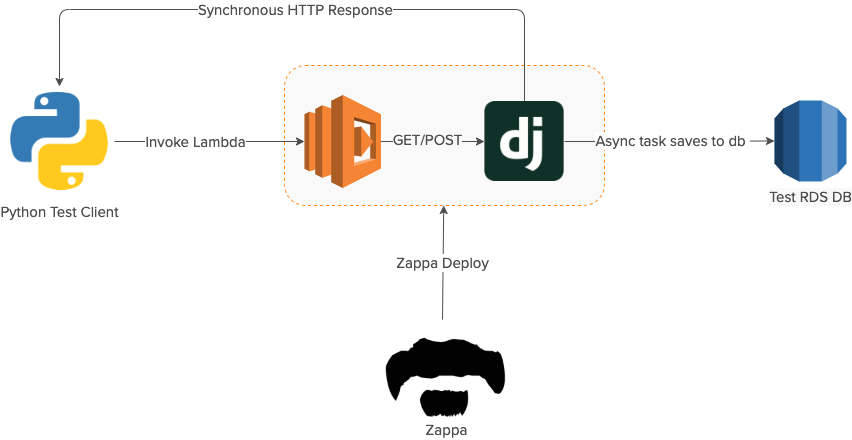
The goal of the prototype was to deploy a Django application to AWS Lambda via Zappa and get that application to run an async task to save data to an RDS database.
The first step was to install and initialize Zappa into a Django application that contains code that can be run asynchronously. A Python virtual environment was executed before Zappa could be executed correctly.
$ pip install zappa $ zappa init
The zappa init command automatically created a zappa_settings.json where we added the necessary configurations for our prototype. Here is what the file ended up looking like:
{
"development": {
"aws_region": "us-east-2",
"apigateway_enabled": false,
"manage_roles": true,
"django_settings": "zappa_prototype.sdk.server.settings",
"project_name": "zappa-prototype-foundry-service",
"runtime": "python3.6",
"s3_bucket": "zappa-prototype-foundry-service",
"timeout_seconds": 900,
"environment_variables": {
"GINKGO_ENVIRONMENT": "development"
},
"vpc_config": {
"SubnetIds": [
"******",
"******"
],
"SecurityGroupIds": [
"******"
]
}
}
}
For the prototype we decided to invoke the lambda function directly rather than use an api gateway. We also gave our AWS account the right privileges so Zappa could easily setup and manage all the roles for the sake of speeding up the prototyping process. In order for Zappa to connect to AWS correctly, the Access Key and Secret Key were set up in ~/.aws/credentials.
[default] aws_access_key_id = XXXXXXXXXXXXXXXXXXXXXXXXXX aws_secret_access_key = XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
After Zappa was configured we added a simple Foundry service asynchronous task to the Django application in the urls.py file. The code accepts either a get or a post request. For the GET request the code responds with “Hello Zappa Prototype”, where as for the POST call the code responds with “Compute DNA Task Received.” HTTP response, while asynchronously starting the 1 minute long compute_dna task. The compute_dna task is analogous with tasks we would have run to process a Foundry service request.
import time
from django.http import HttpResponse
from django.urls import path
from zappa.asynchronous import task
from utils import save_dna_to_test_db
@task
def compute_dna(dna_sequence):
time.sleep(60) # sleep for 1 minute, mimics long task
# Saves dna sequence in the db to acknowledge it was processed
save_dna_to_test_db(dna_sequence)
return "success"
def view(request):
if request.method == "POST":
data = json.loads(request.body)
dna_sequence = data.get("dna_sequence")
compute_dna(dna_sequence)
return HttpResponse("Compute DNA Task Received.")
return HttpResponse("Hello Zappa Prototype!")
urlpatterns = [
path("compute-dna/", view, name="view")
]
After the Django app was all set up we ran the following command to deploy the code into Lambda:
$ zappa deploy development
After the deployment was complete, we ran the following code to verify the Lambda function was running successfully:
import json
import boto3
client = boto3.client("lambda", region_name="us-east-2")
lambda_response = client.invoke(
FunctionName="zappa-prototype-foundry-service",
InvocationType="RequestResponse",
Payload=json.dumps({
"path": "/compute-dna/",
"httpMethod": "GET",
"requestContext": {}
})
)
data = lambda_response["Payload"].read()
lambda_response = client.invoke(
FunctionName="zappa-prototype-foundry-service",
InvocationType="RequestResponse",
Payload=json.dumps({
"path": "/compute-dna/",
"httpMethod": "POST",
"requestContext": {},
"body": json.dumps({"dna_sequence": "ATCGGCTA"})
})
)
data = lambda_response["Payload"].read()
After running this code we successfully got “Hello Zappa Prototype” and “Compute DNA Task Received” in the Lambda responses in a synchronous manner. After a minute, the dna_sequence was successfully saved in the test database, proving the asynchronous tasks ran as expected.
Finally to bring the deployment down we just ran:
$ zappa undeploy development
That’s it! We were able to build a very simple proof of concept to make forward progress with serverless architecture. Zappa gave us capabilities of running 100s of async tasks with very little code.
Whats Next?
The above code is ofcourse a very simplified version of the foundry service and does not capture all the complexities and abstraction around it. We are currently building out an MVP with one of our Foundry services in the serverless architecture using Zappa. Besides providing basic deployment of Django apps, Zappa has been very helpful with its features such as logging and debugging as well as many other advanced customizations.
As we make forward progress it is unclear if we are going to use Zappa outside of MVP, though we are impressed by what Zappa provides. We are committed to the path we have taken towards scaling biology with serverless architectures and will keep you informed of our progress.
